XCP-D : A Robust Postprocessing Pipeline of fMRI data



This fMRI post-processing and noise regression pipeline is developed by the Satterthwaite lab at the University of Pennslyvania (XCP; eXtensible Connectivity Pipeline) and Developmental Cognition and Neuroimaging lab at the University of Minnesota (-DCAN) for open-source software distribution.
About
XCP-D paves the final section of the reproducible and scalable route from the MRI scanner to functional connectivity data in the hands of neuroscientists. We developed XCP-D to extend the BIDS and NiPrep apparatus to the point where data is most commonly consumed and analyzed by neuroscientists studying functional connectivity. Thus, with the development of XCP-D, data can be automatically preprocessed and analyzed in BIDS format, using NiPrep-style containerized code, all the way from the scanner to functional connectivity matrices.
XCP-D picks up right where fMRIprep ends, directly consuming the outputs of fMRIPrep. XCP-D leverages the BIDS and NiPreps frameworks to automatically generate denoised BOLD images, parcellated time series, functional connectivity matrices, and quality assessment reports. XCP-D can also process outputs from: NiBabies, ABCD-BIDS, Minimally preprocessed HCP, and UK Biobank data.
Please note that XCP is only compatible with HCP-YA versions downloaded c.a. Feb 2023 at the moment.
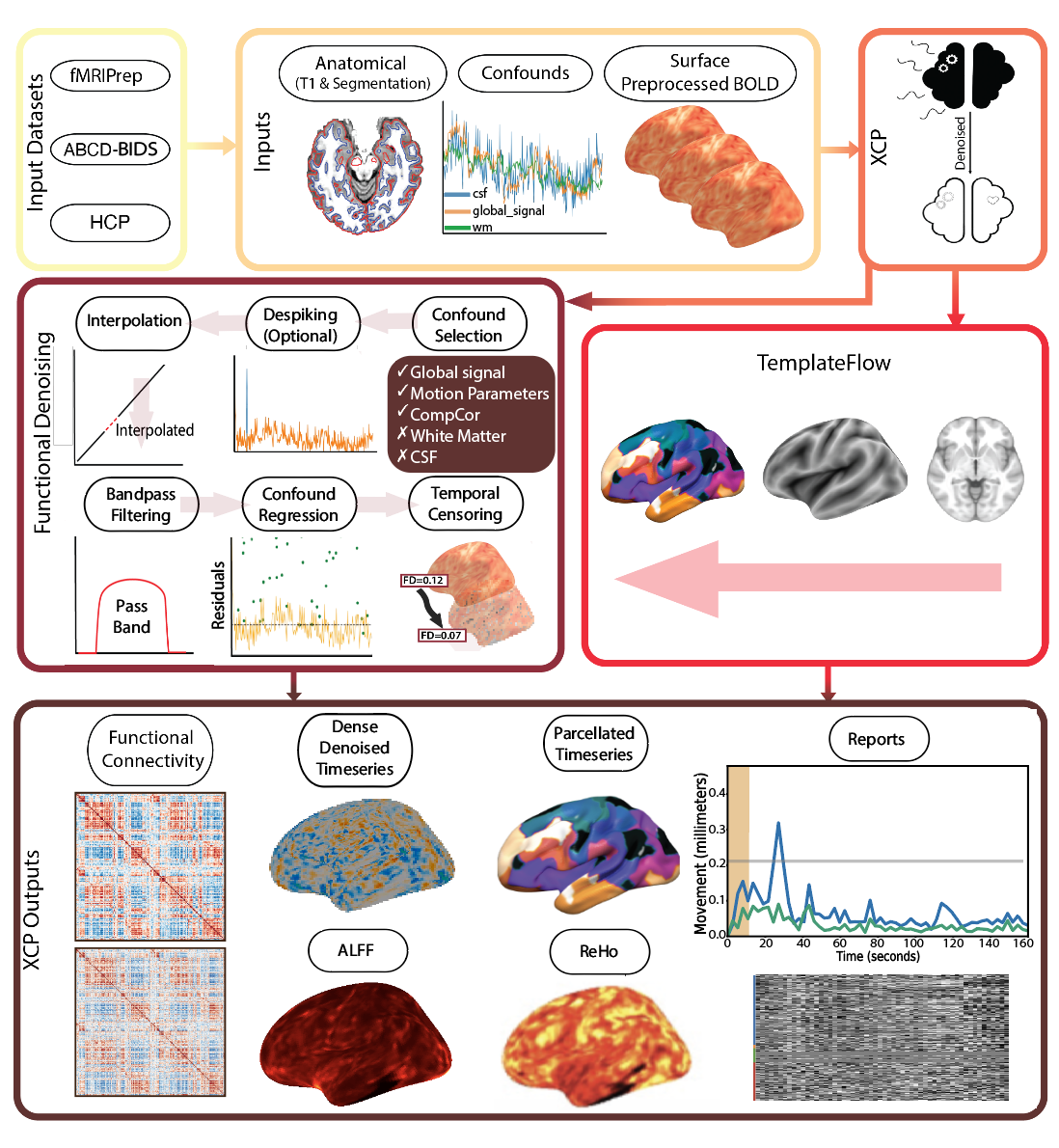
See the documentation for more details.
Why you should use XCP-D
XCP-D produces the following commonly-used outputs: matrices, parcellated time series, dense time series, and additional QC measures.
XCP-D is designed for resting-state or pseudo-resting-state functional connectivity analyses. XCP-D derivatives may be useful for seed-to-voxel and ROI-to-ROI functional connectivity analyses, as well as decomposition-based methods, such as ICA or NMF.
When you should not use XCP-D
XCP-D is not designed as a general-purpose postprocessing pipeline. It is really only appropriate for certain analyses, and other postprocessing/analysis tools are better suited for many types of data/analysis.
XCP-D derivatives are not particularly useful for task-dependent functional connectivity analyses, such as psychophysiological interactions (PPIs) or beta series analyses. It is also not suitable for general task-based analyses, such as standard task GLMs, as we recommend included nuisance regressors in the GLM step, rather than denoising data prior to the GLM.
Citing XCP-D
If you use XCP-D in your research, please use the boilerplate generated by the workflow. If you need an immediate citations, please cite the following preprint:
Mehta, K., Salo, T., Madison, T., Adebimpe, A., Bassett, D. S., Bertolero, M., … & Satterthwaite, T. D. (2023). XCP-D: A Robust Pipeline for the post-processing of fMRI data. bioRxiv. doi:10.1101/2023.11.20.567926.
Please also cite the Zenodo DOI for the version you’re referencing.
License
BSD 3-Clause License
Copyright (c) 2020, Lifespan Informatics and Neuroimaging Center
Redistribution and use in source and binary forms, with or without modification, are permitted provided that the following conditions are met:
Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer.
Redistributions in binary form must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution.
Neither the name of the copyright holder nor the names of its contributors may be used to endorse or promote products derived from this software without specific prior written permission.
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS “AS IS” AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
Contents
- Installation
- Running XCP-D
- Execution and Input Formats
- Command-Line Arguments
- Positional Arguments
- Named Arguments
- Options for filtering BIDS queries
- Options for CIFTI processing
- Options for resource management
- Input flags
- Postprocessing parameters
- Motion filtering parameters
- Censoring and scrubbing options
- Data filtering parameters
- Parcellation options
- Other options
- Experimental options
- Filtering Inputs with BIDS Filter Files
- Running XCP-D via Docker containers
- Running XCP-D via Singularity containers
- Custom Confounds
- Custom Parcellations
- Advanced Applications
- Preprocessing Requirements for XCP-D
- Troubleshooting
- Support and communication
- Processing Pipeline Details
- Input data
- Processing Steps
- Anatomical processing
- Identification of high-motion outlier volumes
- Confound regressor selection
- Dummy scan removal [OPTIONAL]
- Despiking [OPTIONAL]
- Denoising
- Resting-state derivative generation
- Parcellation and functional connectivity estimation [OPTIONAL]
- Smoothing [OPTIONAL]
- Concatenation of functional derivatives [OPTIONAL]
- Quality control
- Outputs
- References
- Outputs of XCP-D
- Contributing to XCP-D
- Developers - API
- What’s New